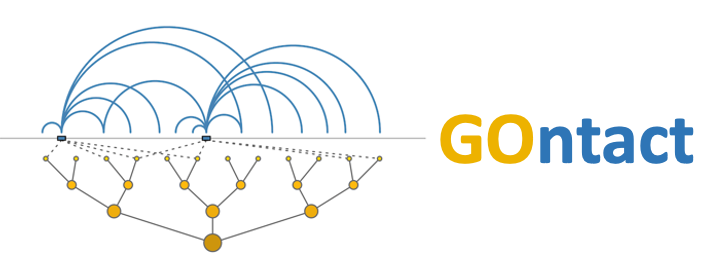
How does GOntact work?
GOntact evaluates Gene Ontology (GO) enrichments for sets of cis-regulatory elements (CREs). To associate CREs to genes, GOntact uses chromatin contacts obtained from chromosome conformation capture data (Hi-C, promoter capture Hi-C, micro-C etc). GO categories are "transmitted" to noncoding elements from the genes that they are in contact with. The statistical significance of GO enrichments is evaluated with binomial tests, which compare the frequency of elements associated with a given GO category in a foreground set (e.g. elements that are specifically active in a given biological condition) with the frequency of elements associated with the same GO category in a background set (e.g. CREs detected in a broader set of biological conditions). If no background set is provided, the expected GO frequencies are computed based on the entire genome.Checkout this page for the source code, a detailed documentation and usage examples for the standalone GOntact version.
Datasets used in the GOntact webserver (version 1.0)
Genome annotations
Genome annotations were downloaded from the Ensembl database, release 115 for human and release 102 for mouse. They correspond to the hg38 and mm10 genome assemblies. These coordinates are used to determine regulatory domains and to annotate baited restriction fragments for the GOntact approach.Gene Ontology annotations
Gene Ontology annotations were downloaded from http://geneontology.org. They correspond to the release 2025-10-10 of the Gene Ontology database. We used the Mouse Genome Informatics (MGI) annotations for mouse and the EBI Gene Ontology Annotation Database for human. We used the "basic" version of the GO database (go-basic.obo) to analyze the hierachical structure of GO annotations and to propagate GO annotations upstream in the graph.Promoter Capture Hi-C data
The data used here were taken from the publication by Laverré et al., Genome Research, 2022. The raw PCHi-C data was combined from multiple sources, listed below. The data was processed with a pipeline based on HiCUP and CHiCAGO.
| species | identifier | tissue/cell type | data source |
| human | NB | naïve B lymphocytes | Javierre et al., 2016 |
| human | TB | total B lymphocytes | Javierre et al., 2016 |
| human | cardio | cardiomyocytes | Choy et al., 2018 |
| human | hESC | embryonic stem cells | Freire-Pritchett et al., 2017 |
| human | EP | endothelial precursors | Javierre et al., 2016 |
| human | Ery | erythroblasts | Javierre et al., 2016 |
| human | FoeT | fetal thymus | Javierre et al., 2016 |
| human | CD34 | CD34+ lymphocytes, hematopoietic progenitors, ex vivo | Mifsud et al., 2015 |
| human | PEK_early | differentiated primary epidermal keratinocytes | Rubin et al., 2017 |
| human | PEK_late | differentiated primary epidermal keratinocytes | Rubin et al., 2017 |
| human | PEK_undiff | primary epidermal keratinocytes | Rubin et al., 2017 |
| human | Bcell | B lymphocytes, EBV-transformed | Mifsud et al., 2015 |
| human | Mac0 | macrophages M0 | Javierre et al., 2016 |
| human | Mac1 | macrophages M1 | Javierre et al., 2016 |
| human | Mac2 | macrophages M2 | Javierre et al., 2016 |
| human | MK | megakaryocytes | Javierre et al., 2016 |
| human | Mon | monocytes | Javierre et al., 2016 |
| human | hNEC | neuroepithelial cells | Freire-Pritchett et al., 2017 |
| human | Neu | neutrophils | Javierre et al., 2016 |
| human | pre_adipo | primary white preadipocytes | Pan et al., 2018 |
| human | NCD4 | naive CD4 T lymphocytes | Javierre et al., 2016 |
| human | NCD8 | naive CD8 T lymphocytes | Javierre et al., 2016 |
| human | TCD4Act | activated total CD4 T lymphocytes | Javierre et al., 2016 |
| human | TCD4MF | total CD4 T lymphocytes | Javierre et al., 2016 |
| human | TCD4Non | non-activated total CD4 T lymphocytes | Javierre et al., 2016 |
| human | TCD8 | total CD8 T lymphocytes | Javierre et al., 2016 |
| mouse | preB_aged | pre-B cells | Koohy et al., 2018 |
| mouse | preB_young | pre-B cells | Koohy et al., 2018 |
| mouse | ESC | embryonic stem cells | Schoenfelder et al., 2015 |
| mouse | ESC_18 | embryonic stem cells | Schoenfelder et al., 2018 |
| mouse | ESC_wild | embryonic stem cells | Novo et al., 2018 |
| mouse | ESC_NKO | embryonic stem cells, Nanog deficient | Novo et al., 2018 |
| mouse | EpiSC | epiblast stem cells | Novo et al., 2018 |
| mouse | ESd_starved | ES-derived multipotent hematopoietic progenitor cells | Comoglio et al., 2018 |
| mouse | ESd_TPO | ES-derived multipotent hematopoietic progenitor cells | Comoglio et al., 2018 |
| mouse | FLC | fetal liver cells | Schoenfelder et al., 2015 |
| mouse | preadip_4H | pre-adipocytes | Siersback et al., 2017 |
| mouse | preadip_D0 | pre-adipocytes | Siersback et al., 2017 |
| mouse | preadip_D2 | pre-adipocytes | Siersback et al., 2017 |
| mouse | TSC | trophoblast stem cells | Schoenfelder et al., 2018 |